GRAS-proteins project
The role of GRAS proteins in developmental biology
GRAS proteins (designated after GAI, RGA and SCR) represent a recently discovered family of plant-specific proteins, whose structure and sequence is highly conserved in the C-terminal part of the proteins (Fig. 1). Although the Arabidopsis genome encodes a family of at least 33 GRAS proteins, only few of them have been characterized regarding their biological function (Fig. 2).
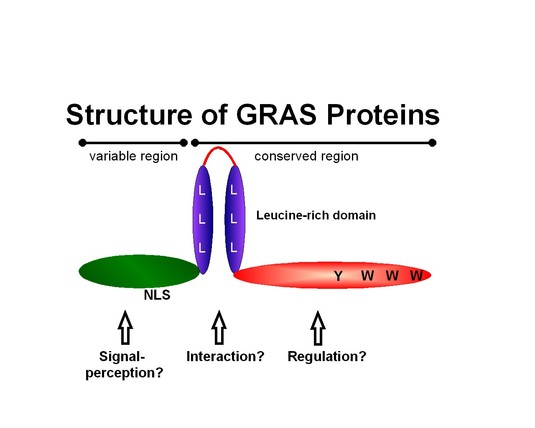
Fig. 1: Schematic organization of GRAS proteins
However, it becomes clear that the functional role of GRAS proteins range from developmental processes such as meristem maintenance to regulation of hormone and light signal transduction. Scarecrow and SHORT-ROOT are involved in root development, HAIRY MERISTEM and LATERAL SUPPRESSOR in meristem differentiation of apical and axillary meristems, respectively. Other GRAS proteins such as GAI and DELLA proteins are involved in gibberellic acid signal transduction, and PAT1 in phytochrome A-dependent signal transduction.
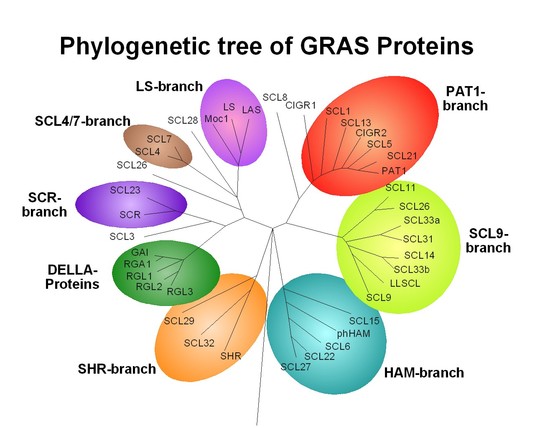
Fig. 2: Phylogenetic tree of the 33 Arabidopsis GRAS proteins including some examples from other plants
The goals for this project are the following:
- The analysis of the biological function of GRAS proteins using a reverse genetics approach and overexpressing lines.
- The cell biological and biochemical characterization of GRAS proteins.
- Structure/function analysis utilizing recombinant DNA-techniques to modify the proteins.
- The identification of interacting proteins, for example with the yeast two-hybrid system.
- The role of specific GRAS proteins for plant architecture and the integration of signals such as light and auxin for the optimized development of plants.
Coworkers:
Publications:
Bolle C. (2004) The role of GRAS proteins in plant signal transduction and development. Planta 218, 683-692
Bolle C. (2009) Phenotyping of Arabidopsis mutants for developmental effects of gene deletions. Methods Mol Biol. 479, 17-34
Bolle C. (2009) Phenotyping of Abiotic Responses and Hormone Treatments in Arabidopsis. Methods Mol Biol. 479, 35-59

